| I wrote Personal Genome Explorer in December 2007 to manage my personal genome data from 23andme. When I originally released it, 23andme didn’t officially support downloading your genome data, so Personal Genome Explorer had to extract the data from the HTML pages from the 23andme website. Now 23andme and deCODEme both officially support downloading your personal genome data, so I’ve deprecated the original HTML crawling path. The latest version imports genome data from either 23andme or deCODEme.
The traits analyzed are taken from SNPedia; if you see any errors, please update the relevant page on SNPedia. Keep in mind that there may be errors in 23andme’s genome data, bugs in how the program imports the data from 23andme, bugs in how the program imports the data from SNPedia, bugs in how the program links the SNPedia info to your genome, or incorrect information on SNPedia. The information presented is meant for recreational purposes only, and I’m not responsible for your interpretation of it. See the license for more details. Features |
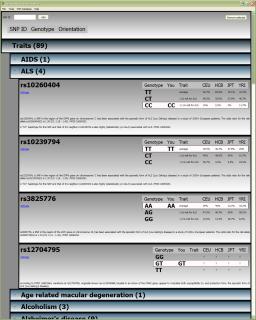 Screenshot Screenshot |
Version History
1.2 (2008-01-27)
- SNPs are grouped by the trait they affect.
- Importing from 23andme‘s official downloadable data format is supported; this has replaced the “screen scraping” data path.
- Importing from deCODEme‘s official download-able data format is supported.
- Genotypes are appropriately oriented on the reference human genome, which makes the cross-referencing between personal genome data and SNPedia more accurate and complete.
- Some miscellaneous UI polish.
1.1 (2008-01-20)
- Initial binary release
- SNP database import from SNPedia
- Comes with pre-imported data from SNPedia
- Analysis of genome based on SNPedia metadata
- Random genome generation based on population frequency data
1.0 (2008-01-12)
- Initial source-code only release
License
You must read and agree to the license before downloading the source or installing the program.
Install
Source download
Version 1.2
The SNP database comes from SNPedia.
It was entirely developed with Visual C# 2008 Express Edition, which is available as a free download.
This project was done as a WPF/.NET learning experience, and I’m happy to receive any constructive criticism you have.
Andrew, I’m astonished! This is a wonderful tool and should be used widely. I wrote a short post about it, I hope you don’t mind.
After several failed attempts at downloading, I got the data and reviewed the traits. the programs seems a little fragile, in that it closes frequently. Perhaps it is just me. But, I saved the data, used a password. Now I cant open the file. I keyed it in the dark, so I could have misstyped. Any suggesttions on how to retrieve the password.
Also data downloaded was primarily for the first three chromosomes. Is 23andme not done testing yet.
Bye the way, thank you very much for the program
Tim, there isn’t a way to open the file without the password; though if you manage to, please let me know, because that would be a bug! 🙂
23andme only started to officially support downloading the raw genome data today, so I will be removing support for downloading the data from their site directly in the next release, and adding support for the officially download-able data.
I’m not aware of any way to crash the application; if you find a way to reproducibly crash it, please post it in the forum and I’ll try to fix it.
Will you consider an extension to include analysis of Affymetrix SNP Array 6.0 data? Should just involve parsing the rs_IDs and passing them into your SNPedia analysis.
This is fantastic, I would like to request an extension to your unbelievable program to include analysis of Affymetrix SNP 6.0 data. This is by far the most widely used platform. Thanks
Sorry guys, I didn’t have time to add support for importing Affymetrix data. I looked briefly for reference material or samples for the format, but I didn’t find anything. Do you have any relevant links?
This looks like a great tool! I have just ordered PG from both 23andme and deCODEme (I am an information junkie!), so I look forward to exploring them using this software.
Just one wish: It would be nice to have a possibility to print the output (or some of it), insteading of just printing a screendump…
affy 6.0 data is supported just fine by promethease
http://www.snpedia.com/index.php?title=Promethease
Pingback: Science Surf » Personal Genome Explorer » Matthias Wjst
Pingback: “Well, Get the Bread Out of the Oven and Let’s Eat!” «
Pingback: Досліди власний геном. « Progenes’s Webblog
Pingback: hongiiv's - 단맛만 좋아요!
Hey, I’m very interested in how you developed this program, and i was wondering, where did you get the data from SNPedia. I couldn’t find such a nice export file.
Hello all, can anyone tell me or direct me to the source regarding how 23andmw or decodeme does data mining? I mean how and what databases they use to compare your or our genotyped data from snp 6.0 arrays? Thanks.
I’m trying to download the personal genome explorer, but get an error message below:
An error occurred trying to download ‘http://www.scheidecker.net/PersonalGenomeExplorer/Personal Genome Explorer.application’.
See the setup log file located at ‘C:\Users\IBM_AD~1\AppData\Local\Temp\VSD147C.tmp\install.log’ for more information.
System.Net.WebException
– The request was aborted: Could not create SSL/TLS secure channel.
– Source: System
– Stack trace:
at System.Net.HttpWebRequest.GetResponse()
at System.Deployment.Application.SystemNetDownloader.DownloadSingleFile(DownloadQueueItem next)